Human-specific genes, cortical development, and brain evolution.
One of the drivers of evolution is the emergence of novel genes through duplication: an ancestral gene is duplicated and the copy evolves into a related, so-called "paralog" gene. Hominid-specific (HS) gene duplications represent a potentially major driving force of evolution, but their impact on human brain evolution remains unclear.
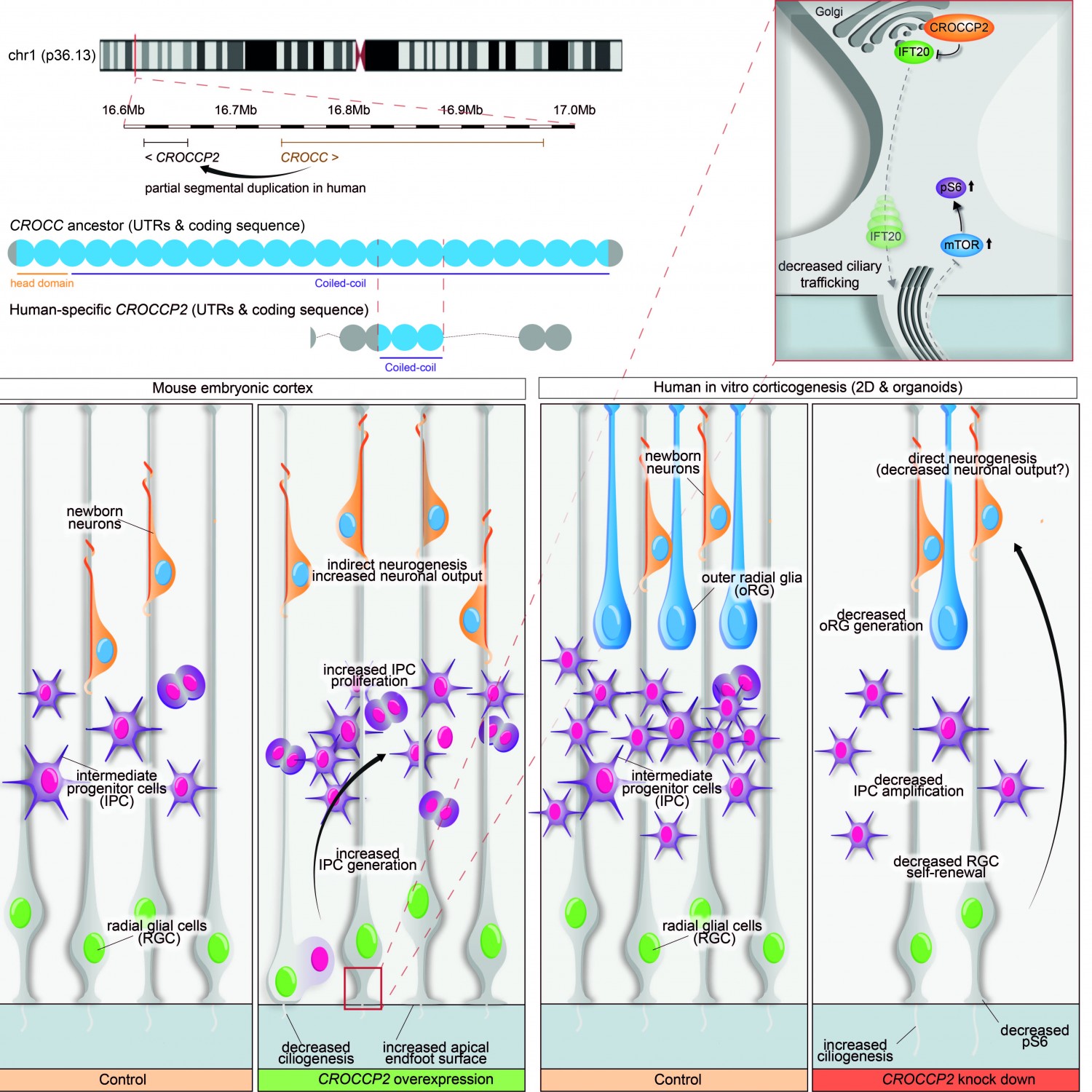
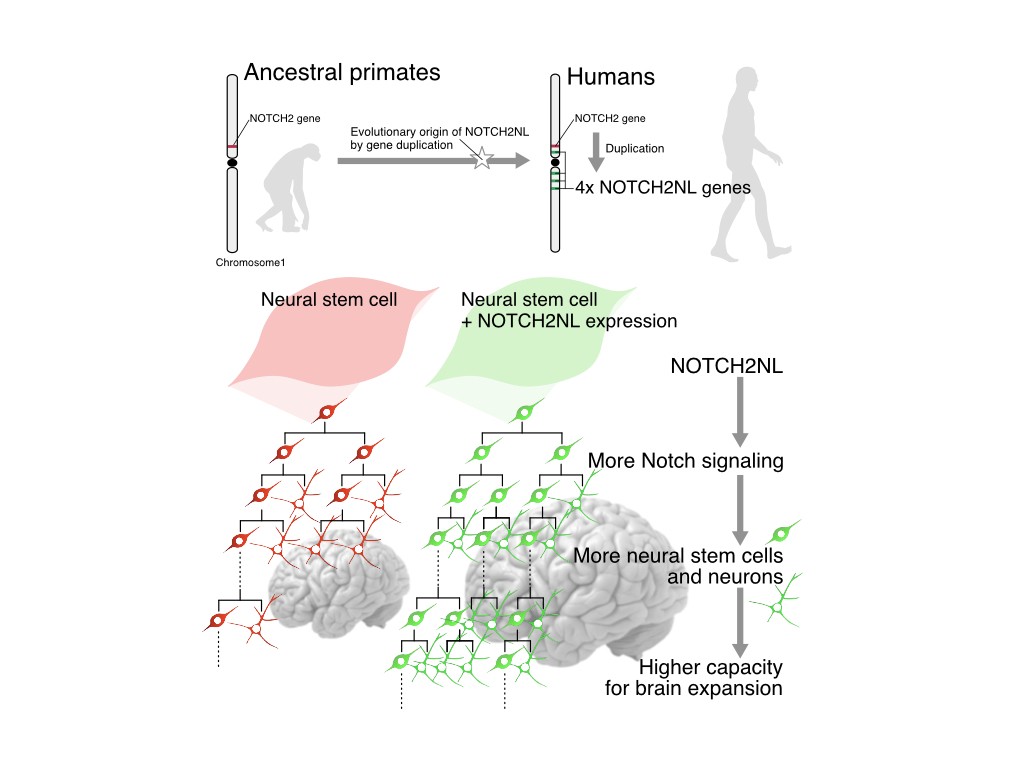
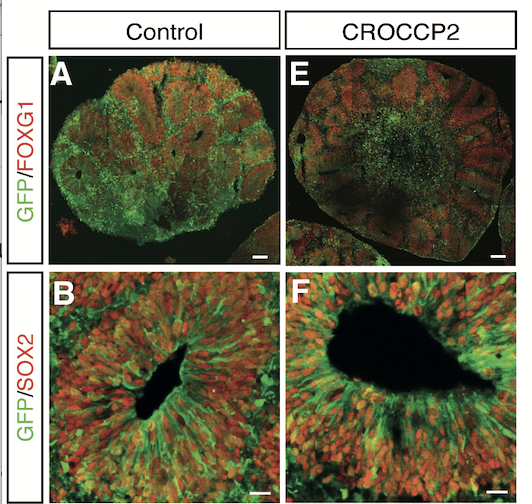
Using tailored RNA sequencing (RNAseq), we recently identified a repertoire of 35 HS genes displaying robust and dynamic patterns during human fetal corticogenesis (Suzuki Cell 2018). Among these we found that NOTCH2NL genes, HS paralogs of NOTCH2, promote the clonal expansion of human cortical progenitors, ultimately leading to higher neuronal production (Suzuki Cell 2018). More recently we identified another HS gene CROCCP2, which contributes to the expansion of human cortical progenitors by modifying ciliary dynamics and enhancing the mTOR pathway (van Heurck Neuron 2023). These data identify HS genes as human-specific modifiers of neurogenesis that may have played an important role in human brain evolutionary expansion.
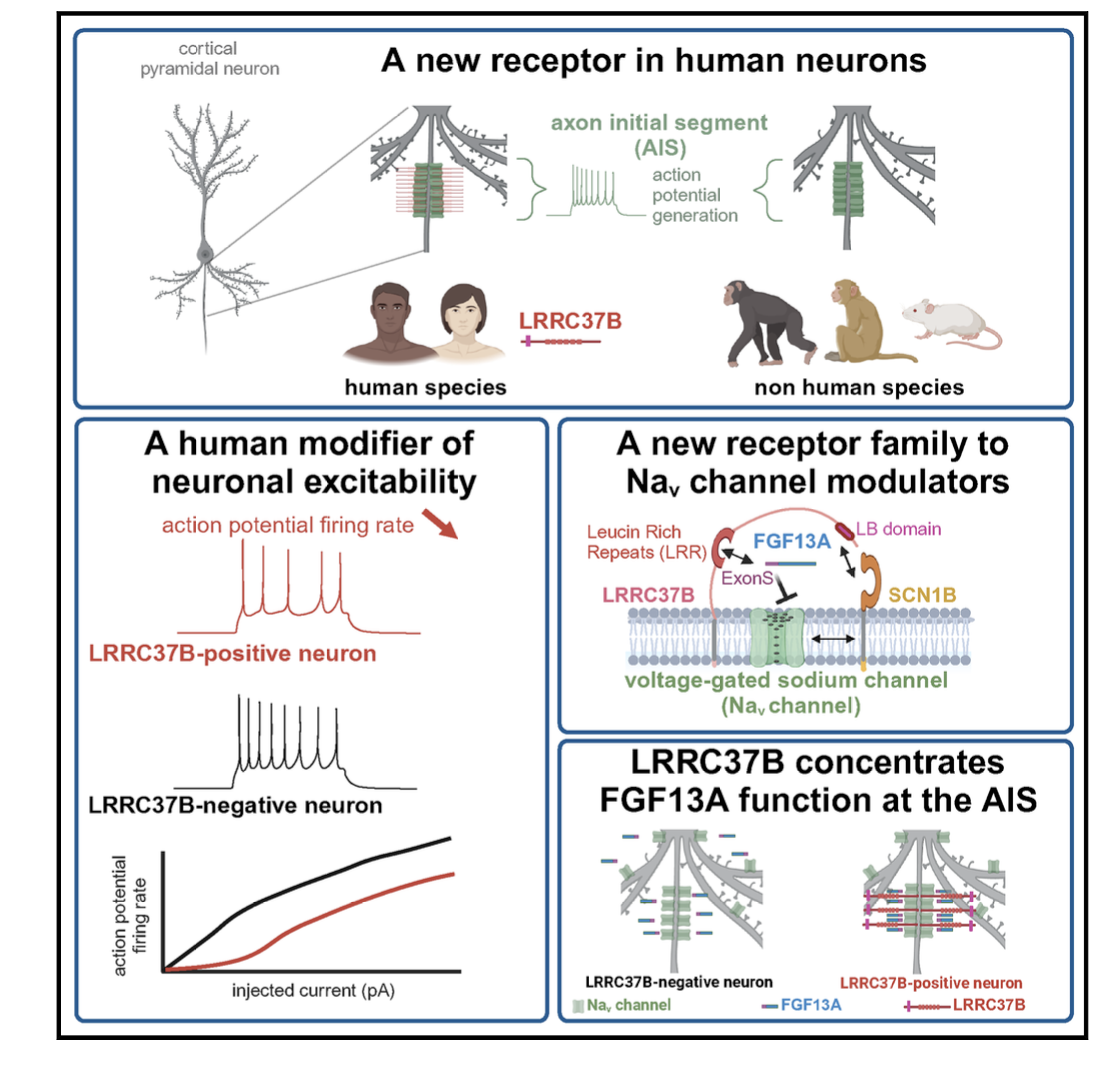
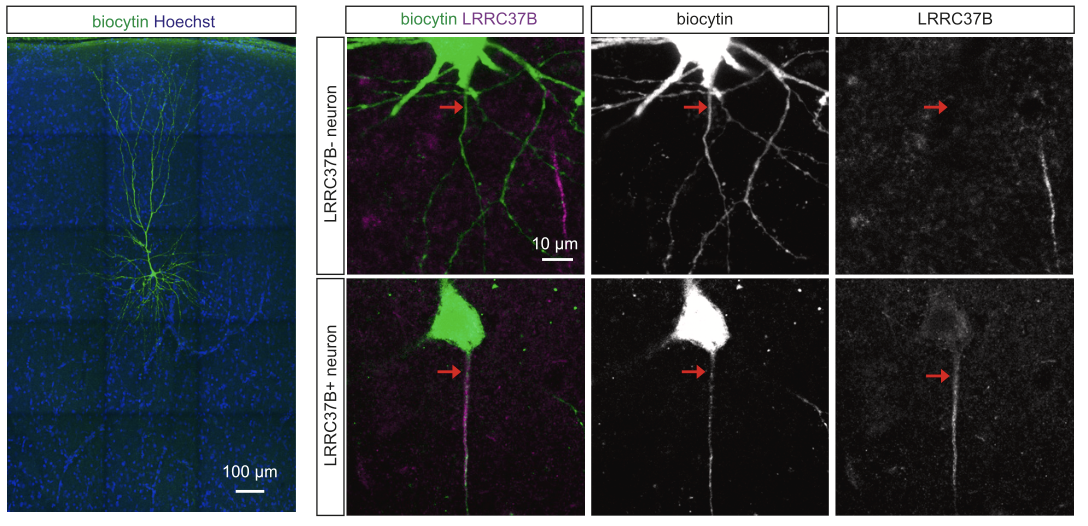
But what about the many other HS genes that we identifed during corticogenesis? Using a combination of gain of function in the mouse, and gain/loss of function in human and non-human primate pluripotent stem cell-based models of corticogenesis, we explore HS gene function from early stages of neurogenesis, to later aspects of neuronal migration, morphogenesis, and synaptogenesis. In particular we recently identified a novel family of receptors, called LRRC37, which are uniquely expressed in human cortical neurons, and control their functional properties (Libé-Philippot et al. Cell 2023). We are now exploring other members of the familar and try to understand how they modify the function of cortical neurons in the context of neural circuits in vivo.